Atomic resolution structure analysis of [4Fe-4S] ferredoxin from B. thermoproteolyticus
Inquiry number
SOL-0000001165
Beamline
BL41XU (Macromolecular Crystallography I)
Scientific keywords
| A. Sample category | atom, molecule, radical, biology, medicine |
|---|---|
| B. Sample category (detail) | atom, biomolecule, crystal, protein, pharmaceuticals |
| C. Technique | X-ray diffraction |
| D. Technique (detail) | single crystal |
| E. Particular condition | low-T (~ liquid N2) |
| F. Photon energy | X-ray (4-40 keV) |
| G. Target information | chemical state, chemical bonding, molecular structure, local structure, structure analysis, charge density |
Industrial keywords
| level 1---Application area | Pharmaceuticals |
|---|---|
| level 2---Target | catalysis, drug design, process analytical technology (PAT) |
| level 3---Target (detail) | protein, drug |
| level 4---Obtainable information | interatomic distance, local structure, electronic state, chemical state, absolute configuration |
| level 5---Technique | X-ray diffraction |
Classification
A80.34 catalysis, A80.50 Pharmaceuticals, M10.10 single crystal diffraction, M10.80 stress strain
Body text
Ferredoxin is a small protein distributing widely in nature, from bacteria to higher plants and animals. It has one or more iron-sulfur cluster, iron atoms binding to thiol base of cysteines in shape of [2Fe-2S], [3Fe-4S] or [4Fe-4S]. The iron-sulfur cluster functions as a redox center for electron transfer, and has other various functions that catalytic site of enzyme, structure stabilizing factor of protein etc.
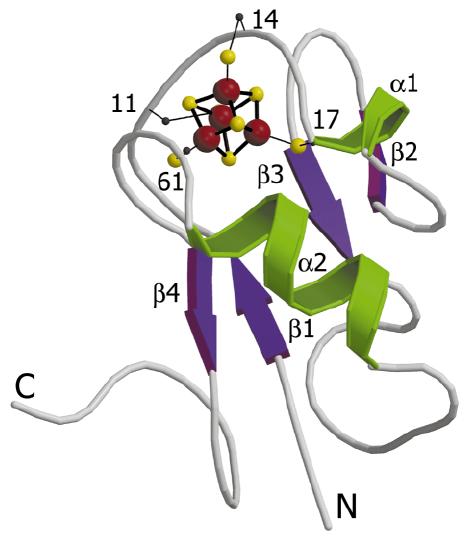
In this case, atomic resolution structure of [4Fe-4S] ferredoxin from B. thermoproteolyticus is analyzed at 0.92 Å resolution (Fig. 1).
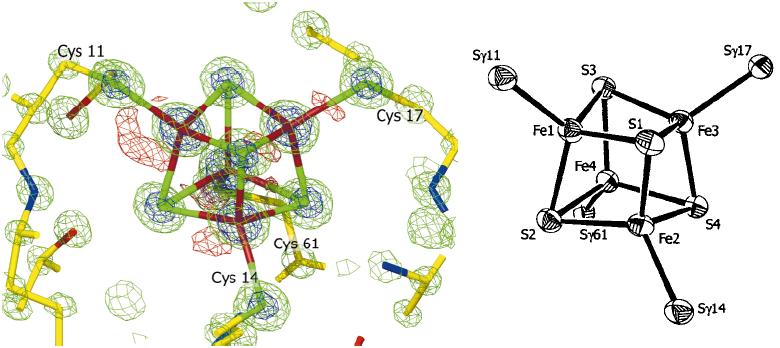
This result revealed that the [4Fe-4S] cluster of this enzyme is distorted (Fig. 2).
Figure 1. Schematic model of [4Fe-4S] ferredoxin from B. thermoproteolyticus.
Figure 2. [4Fe-4S] cluster of electron density map (left) and ORTEP model(right)
[ K. Fukuyama, T. Okada, Y. Kakuta and Y. Yakahashi, Journal of Molecular Biology 315, 1155-1166 (2002), Fig. 1, 4, 7,
©2002 Elsevier, Inc. ]
Source of the figure
Original paper/Journal article
Journal title
K. Fukuyama, et al., J. Mol. Biol., 315, 1155-66 (2002)
Figure No.
1,4,7
Technique
Source of the figure
No figure
Required time for experimental setup
hour(s)
Instruments
| Instrument | Purpose | Performance |
|---|---|---|
| Protein Crystal Diffractometer | To record diffraction data |
References
| Document name |
|---|
| K. Fukuyama, et al., J. Mol. Biol., 315, 1155-66 (2002) |
Related experimental techniques
Questionnaire
This solution is an application of a main instrument of the beamline.
Ease of measurement
Middle
Ease of analysis
With a great skill
How many shifts were needed for taking whole data in the figure?
Two-three shifts