Determination of molecular mechanism underlying peroxisomal disorder rhizomelic chondrodysplasia punctata type 1 (Press Release)
- Release Date
- 01 Jul, 2013
- BL41XU (Structural Biology I)
Key points
• The atomic structure of a peroxisomal protein transport factor in complex with a peroxisomal transport signal sequence has been determined.
• The structure elucidates the mechanism of the recognition of peroxisome targeting signal 2 (PTS2).
• These findings provide the molecular and structural bases for the diagnosis of rhizomelic chondrodysplasia punctata type 1 (RCDP-1) and for the development of new medical treatments of this disease.
|
The research group of Prof. Hiroaki Kato (also, a visiting scientist at RIKEN) and Dr. Dongqing Pan (research scientist, a graduate student at the time of this research), of the Graduate School of Pharmaceutical Sciences, Kyoto University (President, Hiroshi Matsumoto), has determined the molecular mechanism underlying a severe peroxisomal disorder, rhizomelic chondrodysplasia punctata type 1 (RCDP-1)*1. This research was supported by the Targeted Proteins Research Program of the Ministry of Education, Culture, Sports, Science and Technology. Peroxisomes are organelles that play important roles such as catabolism of long-chain fatty acids for energy production. The biogenesis disorder of peroxisomes results in severe peroxisomal diseases. RCDP-1 is caused by the dysfunction of Pex7, a transporter protein that recognizes some indispensable proteins constituting peroxisomes. The mechanism by which Pex7 and a co-receptor protein cooperatively recognize peroxisomal matrix proteins possessing PTS2 was elucidated by determination of the crystal structure of the protein complex containing the three components. The achievements of this group were published online in the scientific journal Nature Structural and Molecular Biology on 30 June 2013 prior to publication in the printed version. |
<<Figures>>
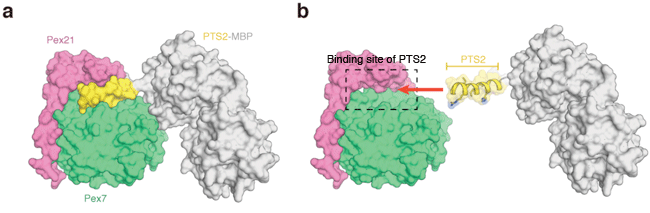
(a) Crystal structures of Pex7, Pex21, and PTS2-MBP (maltose binding protein). PTS2 and MBP are linked by chemical bond and form a protein molecule. The figure shows the surfaces of the three molecules. (b) Binding manner of PTS2 determined on the basis of its crystal structure.
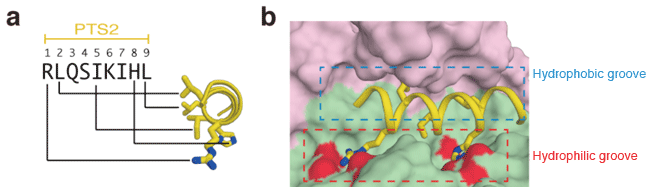
(a) Sequence of nine amino acids of PTS2 and their distribution in the α-helical structure of PTS2. Each letter codes for an amino acid; R for arginine, L for leucine, Q for glutamine, S for serine, I for isoleucine, K for lysine, and H for histidine. The side chains of five important amino acids in the PTS2 helix are shown as a stick model.
(b) Enlarged view of the binding site of PTS2. Pex7 and Pex21 are indicated in green and pink, respectively. The oxygen atoms of acidic amino acids that electrostatically interact with the arginine and histidine of PTS2 are indicated in red.
<<Glossary>>
*1 Rhizomelic chondrodysplasia punctata type 1 (RCDP-1)
RCDP-1 is caused by the dysfunction of Pex7, a protein that recognizes PTS2(peroxisome targeting signal 2), and occurs when the enzymes essential for cellular functions are not transported to peroxisomes. Patients with RCDP-1 show increased levels of phytanic acid and decreased levels of plasmalogen in the body. Patients with severe symptoms often die within a few years of birth, but some patients with a mild dysfunction of Pex7 can live for decades.
|
For more information, please contact: Dr. Dongqing Pan (Kyoto University) |
- Current article
- First observation of electronic structure of carbon-related catalysts during fuel cell operation by soft X-ray emission spectroscopy (Press Release)